|
Research
For a complete list of my publications, please refer to my Google Scholar page.
My current research is in the intersection of accessibility technologies and AI. More specifically, I aim to pose solutions to challenges of people with visual impairments using recent advances in Computer Vision and Machine Learning. The approach I take for doing so is to utilize state-of-the-art deep learning methods that could be used to mitigate the effects of sensory loss experienced by people with visual impairments.
Some of my past research projects are listed below:
Better Segmentation Algorithms for Biomedical Data
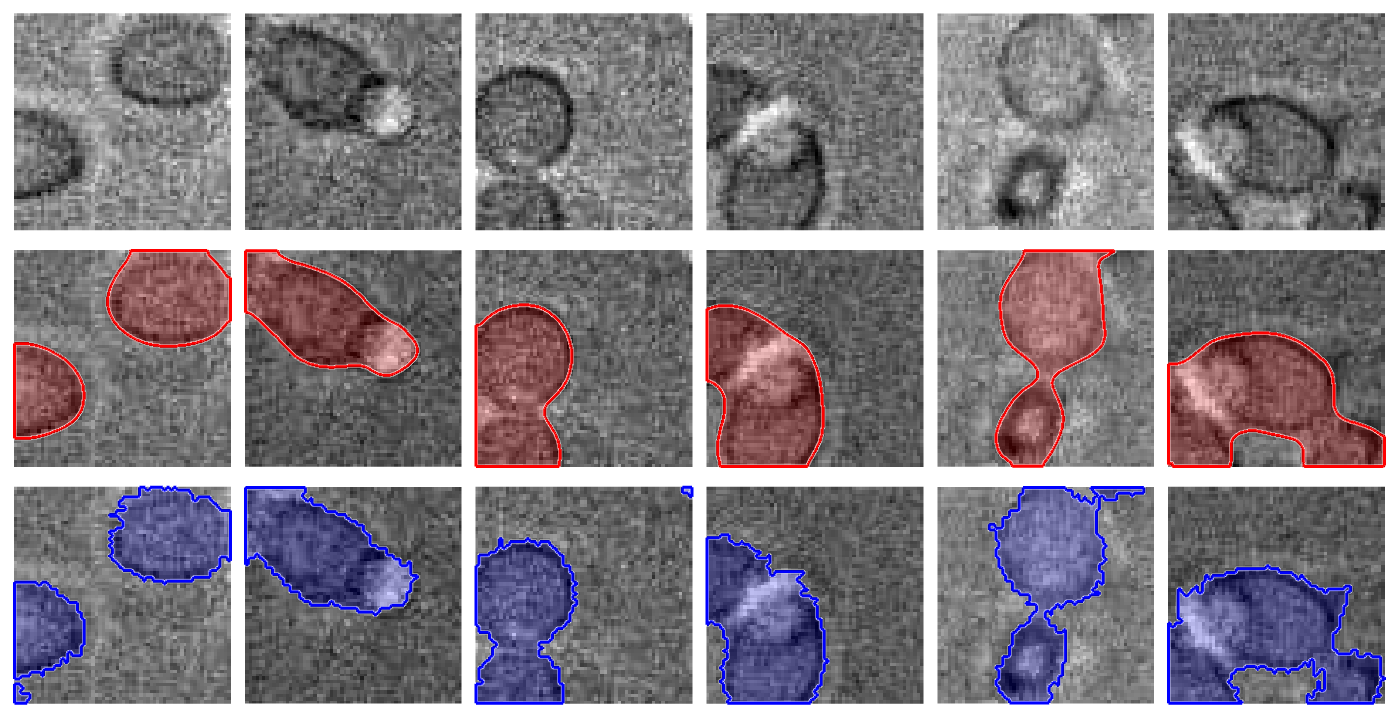 |
One crucial component of quantitative analysis of biomedical data is to develop a robust segmentation solution. While state-of-the-art segmentation algorithms are applicable to this problem, there are challenges and opportunities specific to this type of data. Our goal in this project is to develop segmentation solutions for the analysis of biomedical datasets. In particular, we are interested in analyzing time-lapse microscopy images. Our approach is centered around exploiting multi modal nature of the data. In addition, we treat the sequences of time-lapse microscopy images as video and we attempt to model temporal dependencies across the frames using deep learning architectures. We have published a workshop paper in CVPR2017 on this topic.
|
Efficient Biomedical Dataset Compilation
Gesture(Action) Recognition from 3D Skeleton Data
Prediction of High-Impact Papers Using Temporal and Topological Features
|
